
This is a scientific visualization package for examining output files generated by the Vienna Ab-initio Simulation Package, a package for performing ab-initio quantum-mechanical molecular dynamics using pseudopotentials and a plane wave basis set. The project was initiated when a chemical engineering professor requested assitance in visualizing output files produced by the above package. It displays iso-surfaces and slices of a three-dimensional data set, along with the atoms that make up the molecule the calculations were performed for, and allows symbolic bonds to be inserted between them.
The source code is available for use in non-commerical products via the GNU General Public License. The program was designed to be both reasonably portable and extensible. The only system requirements are an ANSI C compiler, OpenGL 1.1 or later, and GLUT 3.6 or later. The application has been tested under Windows NT 4.0 SP 5 and FreeBSD 3.2, both on Intel processors. Although the program currently only supports the VASP data file format, it should be relatively straightforward to add support for additional file formats.
If you are using a 32-bit Windows system (i.e., Windows 95, Windows 98, Windows NT, or Windows 2000), simply download the Win32 binaries and extract them into a separate directory (a free evaluation version of WinZip can extract the files for you). If you are using a version of Windows 95 prior to OSR 2, you may need to download OpenGL from Microsoft, which is required to run this program.
Once you have extracted the files, and installed OpenGL, if necessary, simply
double-click on vaspview.exe or type vaspview from
the command line, in the directory where you extracted it.
If you are using a UNIX system (such as Linux, FreeBSD, etc.), you will have
to download the source code and compile it for your system. If you download
the .zip file instead of the .tar.gz file, make sure
you use the -a option when extracting the files, to force use of
the correct EOL sequence.
A Makefile is located in the src directory in the source archive.
Edit this Makefile so the CINCLUDE and LIBS variables
refer to the correct locations for include files and library files,
respectively, for your system. Then type
make allto build the executable. It will be placed by default in a
bin
directory in the main vaspview directory.
Type the name of the file you wish to open in the "Open File" text field, and
press <Enter> or click the "Open" button. A progress
indicator (or an error message) will be displayed in the status bar at the
bottom of the window.
Currently, there is no way to browse for a file from within the data viewer. This is because there is no portable way to examine a computer's file system in C.

There are two ways to view portions of the data set: you can view all the data values on a single plane, or slice, that cuts through the data set, or you can view an iso-surface, a surface where all the data has the same value.
The slice may be oriented arbitrarily by changing two angles: the azimuthal angle and the polar angle. These angles define the direction of a vector perpendicular to the plane of the slice. Finally, the slice may be offset along this vector, so that it can be positioned anywhere in the data set.
The controls to manipulate the slice are found on the "Data" tab. You may use
the sliders, or type in a value and press <Enter>.
Alternatively, you can move the slice directly. Click on the slice, or press
<Tab> until it's border is highlighted. Then you can use
the following keys to move the slice:
Left, Right arrowsUp, Down arrowsPage Up, Page DownHomeEndYou can also see the actual data value under the mouse as you move it over the slice, by looking at the read-out at underneath the color legend. The X, Y and Z coordinates listed are the integer coordinates of the actual position in the data set that the mouse is pointing at, and are zero-based.
There are two parameters for the iso-surface. The first is the value of data
the surface should be drawn at. This will create a surface through all the
points in the data set that are equal to this value. Data points that surround
this value will be linearly interpolated to guess the true position of the
surface. Once again, you can specify the value either by moving the slider
control for it, or typing in a value and pressing <Enter>.
The second parameter is the detail level. This controls how much of the data set is sampled when computing the iso-surface, and determines how many polygons will be used to comprise the final surface. The "High" detail level samples every point in the data set, and is as detailed as the surface can get, but this may be very slow on some computers, or with large data sets. The low detail level samples much less of the data set, and should be much faster, though it will miss many of the small and medium features of the surface.

You can also view the atoms that make up the data-set. Each type of atom is
assigned a color, so that all atoms of the same type are the same color. You
may also specify the radius of the atoms (though currently all atoms share the
same radius), by using the slider bar, or typing a number into the "Atom
radius" text field and pressing <Enter>.
You may want to hide some atoms (but not all of them), for a period of time.
There are several ways to do this. You can click on an atom to select it, or
enter it's number in the "Current Atom" text field, and press
<Enter>. You can also use the keyboard, by first clicking
on an atom, or using the <Tab> key until its color changes.
Then the left and right arrow keys will cycle through the atoms, including
those that are hidden (which will temporarily be drawn with a wire-frame
model). Once you have selected an atom, there are several ways to hide it
or make it visible again. You may right-click on the atom, which will
cause it to be replaced by a wire-frame sphere. If you right-click again, it
will become solid again, in case you made a mistake, but if you click
somewhere else, it will completely disappear. You may also use the
<Spacebar> key to toggle whether or not the current
atom is visible. Finally, you can use the "Hide Atom" or "Show Atom" button
on the "Atoms" tab. There is also a "Show All Atoms" button which
will make all hidden atoms visible again.
You may also add a bond from the currently selected atom to another atom.
First select an atom by left-clicking on it, then hold down the
Ctrl key and click on another atom. A bond will now be drawn
between the two atoms. You may also enter the numbers of each atom into the
"Bond from" and "Bond to" text fields, and the desired size of the bond, and
press <Enter> to add the bond. You can change the size of
a bond by selecting it, and then entering the new size in the "Bond size" text
field, or adjusting the slider. You may delete a bond by selecting it, and
pressing the <Backspace> or <Delete>
keys, or with the "Delete Bond" button.
Bonds will be saved in a special file with the same name as the data file you are viewing, except with an ".aux" extension added. You do not need to do anything special to save the bonds, it is done automatically as you edit them.
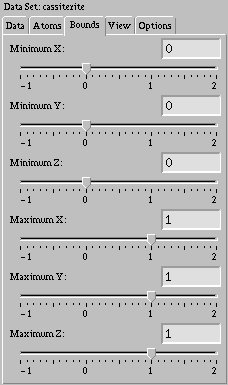
You can change the amount of the data set being viewed by changing the size of the box surrounding it. If you make the box smaller, you can zoom in to particular parts of the data without having to worry about the parts around it. If you make the box larger, the data set will be repeated so you can see how molecules with periodic boundary conditions appear.
You may change the position of any one of the sides of the box by moving the
appropriate slider in the "Bounds" tab, or by typing in a value and pressing
<Enter>. The default minimum value is zero, and the default
maximum value is one. Be warned that if you make the box very large,
performance may degrade from having to draw the data set up to twenty-seven
times over.
You can also change the size of the box with the keyboard. Press the
<Tab> key until one of the sides of the box is highlighted
in green, and then the following keys may be used to control the size of the
box:
Left, Right arrowsUp, Down arrows, Page Up, Page DownHomeEndFinally, you can left-click somewhere on the background of the viewport, and drag out a rectangle that will clip the box. While you are dragging, the actual portion of the box you are selecting will be highlighted in green. If you make a mistake, or want to back up, you can right-click on the background to restore your previous view.
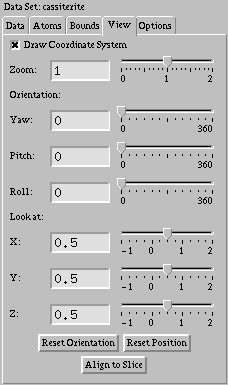

There are several parameters which control how you are looking at the data
set. The first is the zoom factor. The smaller this number, the closer you are
to the point you are looking at. The larger, the farther away. You can set
this by using the "Zoom" slider on the "View" tab, or by typing a number into
the text field and pressing <Enter>.
The next set of parameters control the orientation of the data set. These are
three angles: yaw, pitch and roll. Each one controls a rotation about a
different axis, and together they can specify any orientation for the data
set. You can change them by using the sliders on the "View" tab, or by typing
in a number in their text fields, and pressing <Enter>.
However, it is usually much more convenient to drag the data set into the
orientation you want. You can do this by moving your mouse over one of the
edges of the box, or one of the coordinate axes, and clicking and dragging.
The data set will be rotated around the point you are looking at so that the
point you clicked on is always under your mouse.
The final set of parameters is the point you are looking at. By default, this
is the center of the data set. The point you are looking at appears in the
center of the viewport, and is the point the data set is rotated around when
you change the angles above. You can change this by using the sliders on the
"View" tab, or by typing in a number into the corresponding text fields, and
pressing <Enter>.
Finally, there are three special buttons provided on the "View" tab. The first, "Reset Orientation," resets all three angles that control your orientation to zero. The second, "Reset Position," will move the point you are looking at back to the center of the data set. The third, "Align to Slice," will change your orientation so that you are looking directly at the slice.
All of these things may be controlled directly with the keyboard as well.
Either click on the background of the viewport, or press
<Tab> until the coordinate axes are highlighted in green,
and then you can use the following keys to control your view:
Page Up, Page DownLeft, Right arrowsUp, Down arrowsShift + Left, Right arrowsHomeEndCtrl + Left, Right arrowsCtrl + Up, Down arrowsCtrl + Page Up, Page DownCtrl + Home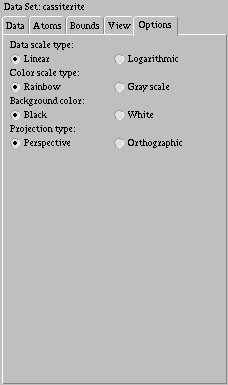
There are several other options you can use to control how you view the data
set. The first is the "Data scale type," which controls how data values are
mapped into colors. There are two options, "Linear" and "Logarithmic." A
linear map simply scales the data into the range 0 to 1, and assigns a color
to each value in that range. The logarithmic map scales the data linearly into
the range 1 to e2. Then the natural log of the
resulting value is taken, and divided by 2, to provide a number between 0 and
1, which is then assigned a color.
The next option, "Color scale type," determines what colors are used to display the data values on the slice and to distinguish different types of atoms. There are two options, "Rainbow" and "Grayscale." The rainbow scale goes from blue for small values, through green, and ends up with red at large values. The gray scale goes from black for low values to white for high values. While a rainbow scale is often clearer, the gray scale may be better for printing.
The next option, "Background color," controls what color is used as a backdrop for the data set. There are two choices, white and black. Black often provides better contrast, but white may be useful for printing.
The final option is the "Projection type." This controls whether or not the data set is shown with 3D perspective, so you can judge depth information, or if it is projected straight onto viewport (orthographic). While the first often gives you a better idea of the relative positions of objects, the second is useful when looking along an axis to see how things line up.
There no facility provided within this program to save or print images of a
data set. However, your computer may already provide you with the tools to do
this. If you are running Windows, you can use <Alt> +
<PrintScreen> to copy the contents of the current window to the
clipboard. You can then use one of the utilities provided in your
"Accessories" menu off your start bar to paste the window into an image to be
saved or printed. If you are using UNIX, the popular program
XV offers similar functionality, and is
shareware.
This program can only read files ouput by VASP version 4.4.4 or later. Earlier versions used fixed-format FORTRAN files, while later versions used free-format, which is incompatible.
This program was written primarily for Windows, though it can be compiled and
run anywhere that supports OpenGL and the GL Utility Toolkit. As such, it can
only use the features of OpenGL 1.1, since this is the latest version shipping
with Windows as of the time of this writing. OpenGL 1.2 provides an additional
feature known as 3D texturing, which substantially improves the speed and
quality of the rendered slice. The data viewer can take advantage of this
feature, but only if it is compiled on a system that supports it. The Win32
binaries distributed here have not been compiled on such a system, and so if
you want to use it, you will have to recompile the program yourself. Some
systems claim to support 3D texturing, but I have been unable to get it to work
on them (some Linux machines and some SGI machines). If you experience this
problem, add the switch -D__DS3_NO_3D_TEXTURES__ to the
CFLAGS variable in the Makefile under UNIX, or add
__DS3_NO_3D_TEXTURES__ to the list at "Project / Settings /
C/C++ / General / Preprocessor definitions" under Windows.
The <Delete> key does not work under Windows. This is a bug
in the GL Utility Toolkit that will hopefully be addressed in future versions.
You can use the <Backspace> key instead.
The Shift + <Tab> key does not work under some
versions X-Windows. This is a bug in the GL Utility Toolkit that will hopefully
be addressed in future versions. This will prevent you from changing the
keyboard focus from component to component in the reverse order. You can use
any other modifier key (e.g. Ctrl or
Alt) instead.
I would like to offer special thanks to Dr. David Cox, in Chemical Engineering at Virginia Tech. He gave me some valuable advice when trying out the alpha versions of the viewer. I would also like to thank Dr. Doug Bowman, in Computer Science at Virginia Tech, who gave me a few good ideas for the direct manipulation of the data set.
Binaries are availble for 32-bit Windows systems. The source code is available under the GNU General Public License, and includes an MSVC++ 5 project file and a Makefile (that may require manual editing; if you want to write a more general Makefile, or just one for a specific platform, feel free to send it to me). If you use this program, or make modifications to the source, please drop me a line.
| Comments or questsions? Send them to tterribe@users.sourceforge.net |
|